Mapping with R2RML
It is not uncommon when accessing virtualized database tables that the data stored in them is not particularly well-structured for direct use as semantic data. Database columns may be incorrectly or obscurely named, resulting in obscure IRIs. They may have data types different from what a well designed semantic data source would. Data may be overly normalized, resulting in lots of extra triples in the graph and overly verbose queries against it. To help with all of these data presentation issues, data.world supports extending your datasets with predicates and classes defined via the R2RML ontology.
R2RML allows for predicates and classes to be defined by reference to relational tables and columns, or by SQL queries over those tables and columns. With R2RML, you can create semantically meaningful SQL queries defining classes and predicates and then you can use those classes and predicates in SPARQL queries on data.world.
To apply an R2RML mapping to a data.world dataset:
create an R2RML mapping file that refers to the tables in that dataset
upload that file to the dataset.
To access the classes and predicates you have defined via R2RML, use the :mapped named graph, just as with virtualized tables. Further information about the R2RML ontology and how it allows predicates and classes to defined can be found here.
Example - R2RML
Important
All the links in the following examples open datasets/projects in the data.world open data community. For best experience, make sure you are logged in to the application when you click these links.
Here is a sample query using R2RML:
PREFIX : <https://ddw-doccorp.linked.data.world/d/sparql-r2rml/>
SELECT ?abbreviation ?normalized_abbreviation ?runways
{ SERVICE :mapped {
?s <http://ddw-doccorp.linked.data.world/d/sparql-r2rml/col-extended_airports-abbreviation> ?abbreviation.
?s <http://ddw-doccorp.linked.data.world/d/sparql-r2rml/col-extended_airports-normalized_abbreviation> ?normalized_abbreviation.
?s <http://ddw-doccorp.linked.data.world/d/sparql-r2rml/col-extended_airports-runways> ?runways.
}
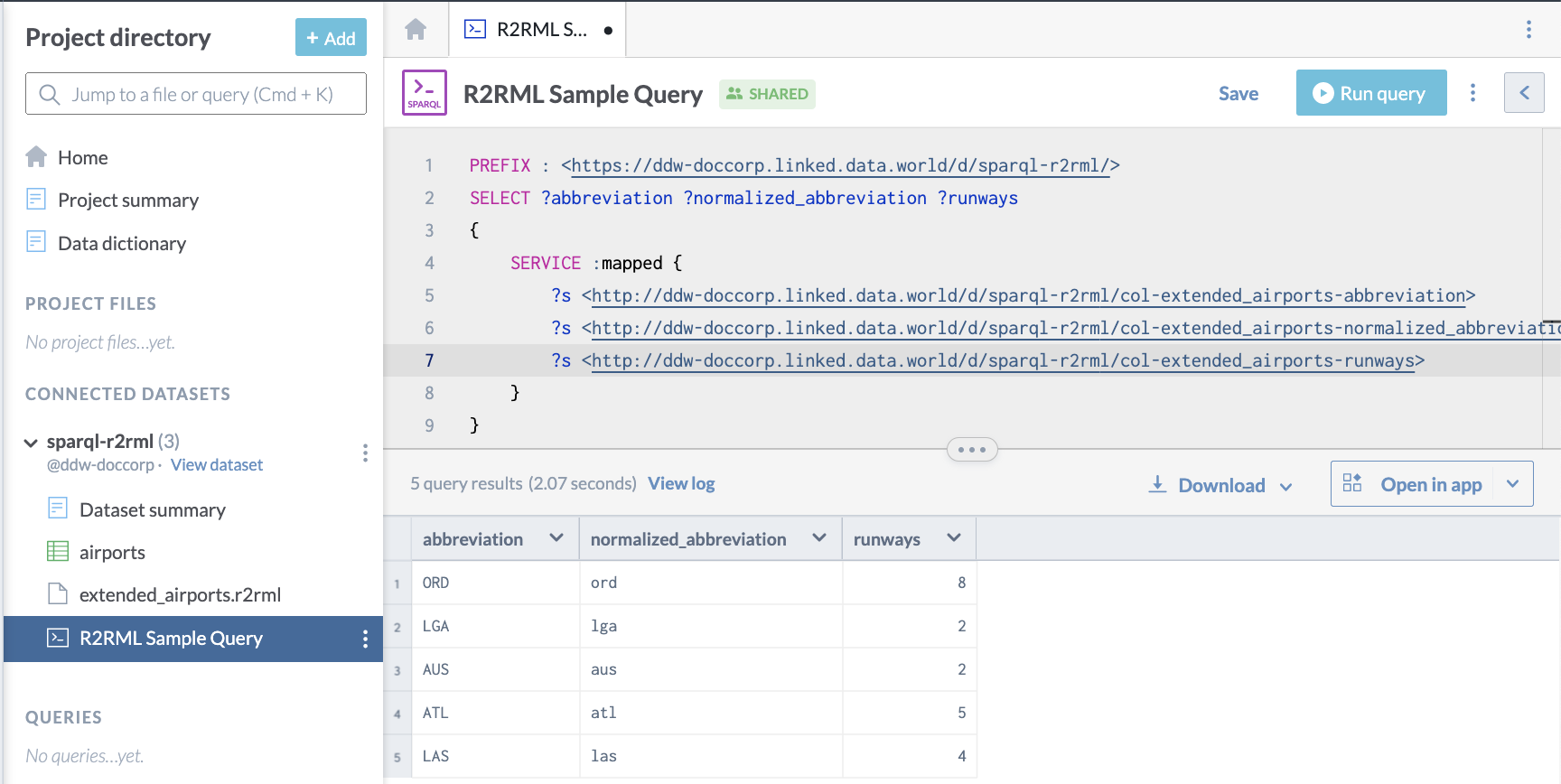
}This is what it looks like on data.world: